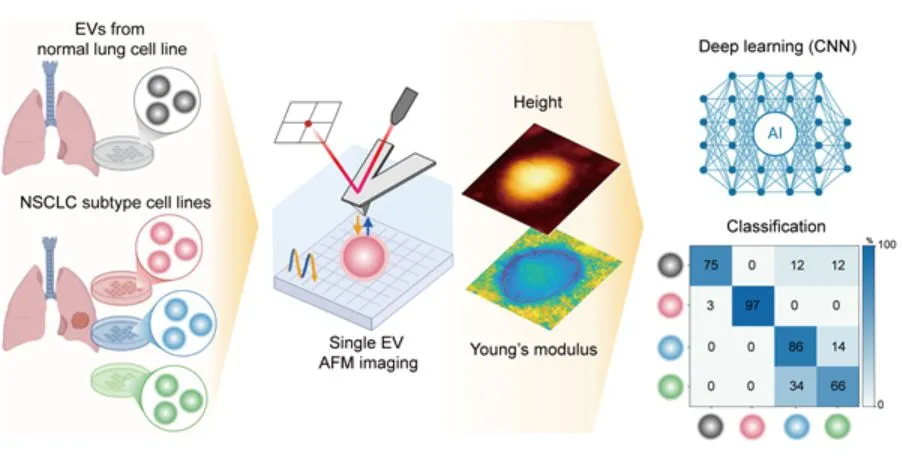
DGIST researchers developed a deep learning model that classifies lung cancer exosomes based on physical properties measured by atomic force microscopy.
Key Details
- 1DGIST team used AFM to measure nanomechanical properties (stiffness, height-to-radius) of exosomes from NSCLC cell lines with different genetic mutations.
- 2AI model (DenseNet-121) classified exosomes by origin, achieving 96% accuracy and AUC of 0.92.
- 3Exosome stiffness reflected KRAS and EGFR mutations in their respective lung cancer cell lines.
- 4The method enables high-precision, label-free, liquid biopsy-based lung cancer diagnosis.
- 5Study published July 8, 2025, in Analytical Chemistry.
Why It Matters

Source
EurekAlert
Related News

Researchers Develop All-Optical Synapse for Neuromorphic Imaging Systems
A new artificial synapse, controlled entirely by light, enables in-sensor neuromorphic processing for more efficient and noise-resistant imaging systems.

AI-Simulation Approach Achieves 90% Faster Brain MRI with Minimal Data
A simulation-based AI method can reconstruct brain MRI scans with only 10% of the usual data, greatly reducing scan times.

Mayo Clinic Showcases Imaging AI and Early Cancer Detection Advances at ASCO 2026
Mayo Clinic researchers will present over 30 studies at ASCO 2026, highlighting new advances in imaging AI, data science, and early cancer detection.