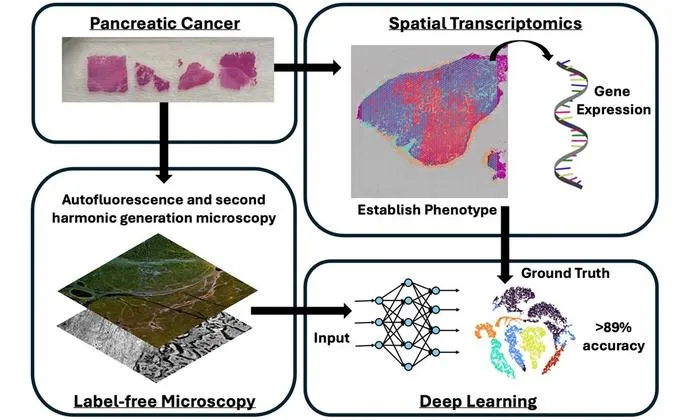
University of Arizona researchers achieved nearly 90% accuracy in pancreatic cancer phenotyping using label-free optical microscopy with deep learning AI.
Key Details
- 1Label-free optical microscopy paired with deep neural networks identified tissue phenotypes at over 89% accuracy in pancreatic cancer samples.
- 2Spatial transcriptomics served as the 'ground truth' for phenotypic classification.
- 3Traditional image analysis could not match the performance of AI methods, pointing to AI's necessity in extracting meaningful features from label-free images.
- 4This approach bypasses expensive and time-intensive molecular/genetic sequencing currently used in precision medicine.
- 5The work demonstrates a significant step toward more accessible and rapid phenotyping for cancer care.
Why It Matters

Source
EurekAlert
Related News

Researchers Develop All-Optical Synapse for Neuromorphic Imaging Systems
A new artificial synapse, controlled entirely by light, enables in-sensor neuromorphic processing for more efficient and noise-resistant imaging systems.

AI-Simulation Approach Achieves 90% Faster Brain MRI with Minimal Data
A simulation-based AI method can reconstruct brain MRI scans with only 10% of the usual data, greatly reducing scan times.

Mayo Clinic Showcases Imaging AI and Early Cancer Detection Advances at ASCO 2026
Mayo Clinic researchers will present over 30 studies at ASCO 2026, highlighting new advances in imaging AI, data science, and early cancer detection.